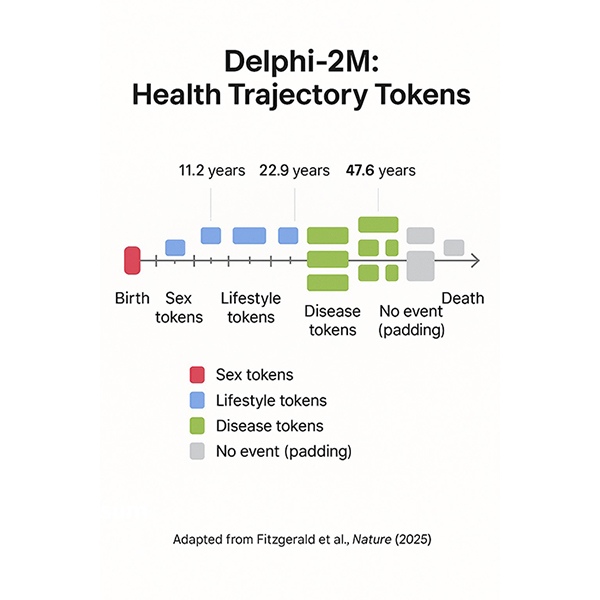
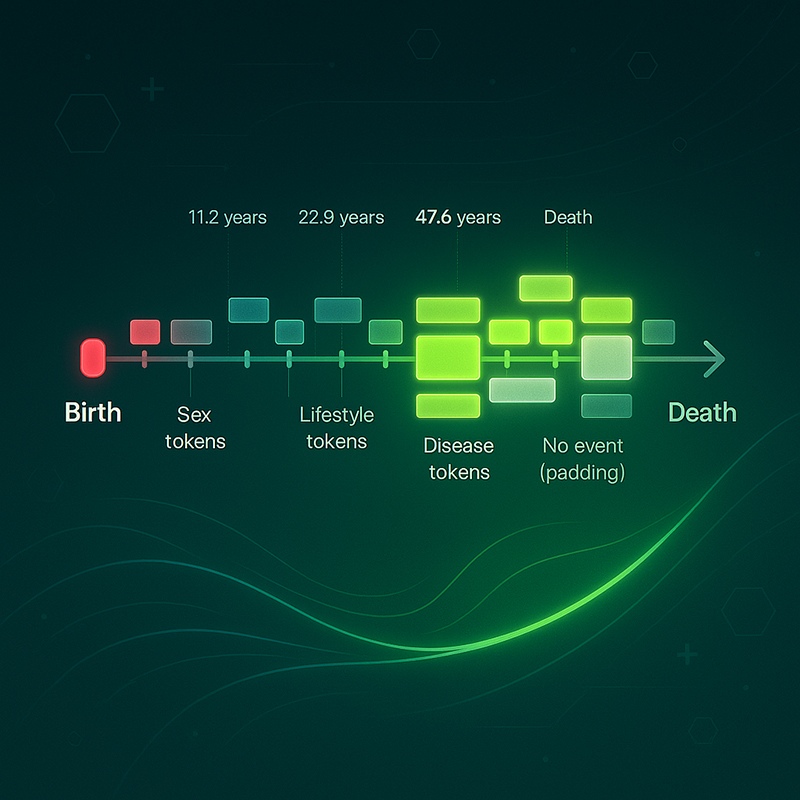
A generative model estimates risks for 1,000+ conditions and times those risks like a weather forecast. If validated and governed well, it could shift care from reaction to prevention.
Preventive medicine is edging from hindsight toward foresight. A new artificial intelligence system, Delphi 2M, created by teams at EMBL’s European Bioinformatics Institute, the German Cancer Research Centre, and the University of Copenhagen, forecasts a person’s risks across more than one thousand conditions and places those probabilities on a timeline years into the future.
Trained on anonymized records from 400,000 UK Biobank participants and 1.9 million patients in Denmark, the model learns patterns in medical events and lifestyle factors, then expresses risk as rates over time, much like a weather forecast. Findings published in Nature on September 17, 2025, report accuracy comparable to leading single disease tools, with the added ability to simulate future health pathways and account for competing risks and multimorbidity. We spoke with Tom Fitzgerald, a senior staff scientist at EMBL EBI, about what this could mean in practice. Quotes have been edited for length and clarity.
Beyond single scores: modeling whole health trajectories
“Historically, it has been difficult to represent large numbers of individuals across a very large disease spectrum and across time,” Fitzgerald says. Where classical approaches build one model per disease, Delphi-2M considers many at once and maps risk to time. Key innovations include an explicit time-encoding scheme that lets the team optimize both what comes next and when it is likely to happen, with calibration that can be checked at future time points. “Given any starting point in someone’s life, we can generate likely trajectories years ahead,” he adds. For some conditions, performance remains accurate for more than a decade. The system is generative, so it can sample synthetic future trajectories to estimate potential disease burden over one to two decades.
The model produces probabilities, not certainties, which is why the weather analogy fits. Short-horizon forecasts often work best for acute conditions, while longer horizons can favor chronic diseases. In addition to point estimates, risk can be accompanied by uncertainty bounds for a person’s age bracket and condition, which helps communicate confidence to clinicians and patients.
Data in, data out: demographics, genetics, and the environment
The published model uses health history and a small set of demographics: smoking status, alcohol consumption, body mass index, age, and sex. Race is not an input feature in this first release. Genetics is also not included in the paper’s main model, although the team has prototypes with polygenic risk scores and HLA alleles that improve performance for some diseases and can help separate distinct biological pathways in conditions such as diabetes. There are open technical questions about how best to represent the genome inside these models, and the base system is already very performant without it. The team also sees value in adding environmental and geographic signals, for example pollution levels, to improve prediction and to study how risks change with social or environmental conditions.
Performance patterns, equity, and rare disease
Counterintuitively, the model can perform better for some minority ancestry groups and for people in more socially deprived settings than for the majority white UK cohort. Fitzgerald links this to data density: where disease is more common, there is more signal to learn from. This does not imply better care. It underscores the need for subgroup audits, local validation, and recalibration to ensure fair use across populations. Sparse health records can reduce performance for anyone, including majority groups.
Rare diseases remain harder. Performance improves with larger datasets and richer histories. The current model uses ICD-10 codes at a level that balances specificity with data sufficiency. With larger training cohorts, the team could model more specific code levels, which may help for very rare conditions. Value may also come from earlier warnings that shorten diagnostic odysseys rather than from headline accuracy numbers alone.
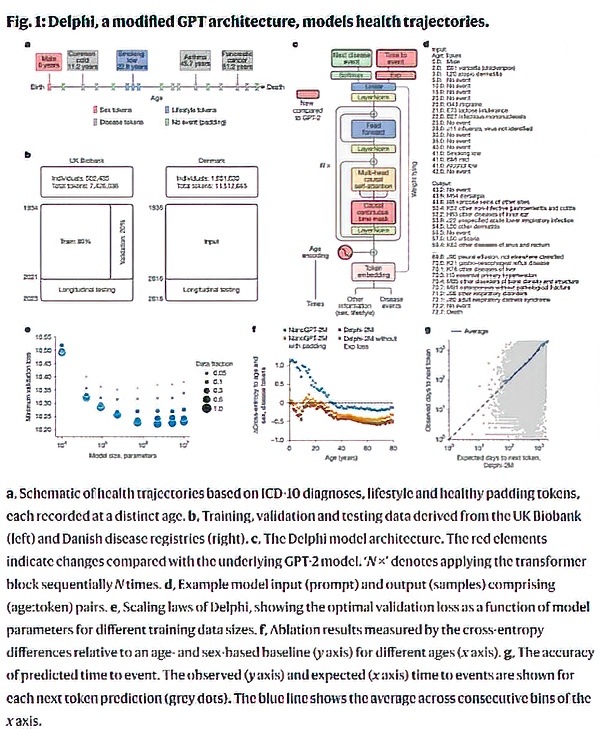
From lab to clinic
Delphi-2M is not a clinical tool yet. “There needs to be more testing, evaluation, and policy review before clinical applications,” Fitzgerald says. The near-term promise lies in early-warning workflows that flag rising risks over the next one to two years and in resource planning that helps systems anticipate demand across regions and providers. EMBL interim executive director Ewan Birney has suggested patients could benefit within a few years. Fitzgerald adds that individual-level early warning may be five to ten years away, since it will require prospective trials, external validation, careful presentation of uncertainty, and clinician-friendly interfaces. The group already collaborates with clinicians in a research capacity to understand what information is most useful.
Governance, portability, and drift
The model was trained in UK Biobank, evaluated on a held-out UK set, then transferred to Denmark with only a marginal performance drop, which suggests it learned patterns that generalize across systems. Even so, any use elsewhere requires local validation and calibration. Derived model weights from the UK training were returned to UK Biobank and are accessed under that program’s governance rules. Predictions for individuals require individual-level data, which would remain under the control of the local institution. Re-identification from model weights alone is considered unlikely, though not impossible, so governance proceeds with caution.
Real deployments must plan for distribution shift as coding practices, populations, and policies change. The team has not yet run a dedicated longitudinal drift study. COVID is an obvious case study that could reveal how delivery and disease rates before and after the pandemic differ and how models should adapt. Any clinical use will need monitoring, rollback plans, and human-in-the-loop safeguards.
Why a generative approach
Generative modeling enables the system to represent competing risks and multimorbidity across time, which is difficult with classical per-disease models such as Cox regressions. This unified approach helps align large numbers of people across time and disease state and supports both individual-level and population-level questions, from likely next events to multi-year resource planning.
Who built it, and how long did it take
Fitzgerald is a senior staff scientist at EMBL-EBI with more than twenty years in large-scale cohort analysis and genomics. He previously worked at the Wellcome Sanger Institute on the Deciphering Developmental Disorders project, the UK’s first very large sequencing effort with direct clinical application for rare disease, and for the last decade has co-run Ewan Birney’s research group at EMBL-EBI. Delphi-2M took roughly a year and a half from inception to a functioning system and about two years to publication, with contributions from multiple teams in Cambridge and Heidelberg.
What comes next
The team is improving the model by adding more data modalities, including genetics and environmental context, and by studying how best to present risk and uncertainty in ways clinicians can use. Success will be measured by whether acting on forecasts improves outcomes, reduces inequities, and avoids harm.
Conclusion
Delphi-2M marks a notable step for predictive medicine: one model that times risk for many conditions and sketches the likely path ahead. The Nature results across UK and Danish data are strong. Clinical use, however, hinges on prospective trials, fairness audits, governance, and careful integration into workflows. The technical capability to look years ahead now exists. The task is to ensure that foresight is useful, fair, and safe for the people it aims to serve.
Disclosure: Quotes have been edited for length and clarity. This article is for information only and is not medical advice.